|
4/26/2023 0 Comments Numpy vstack names array columns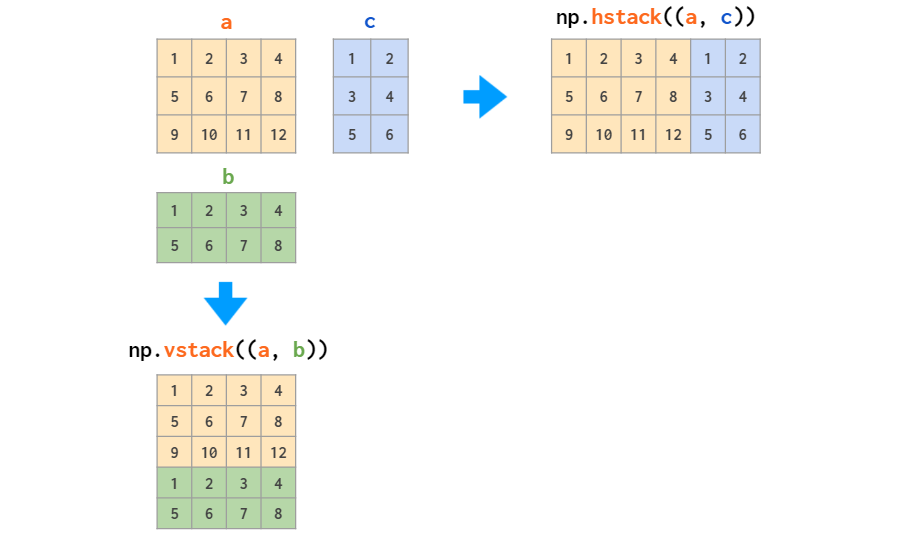
The array formed by stacking the given arrays.Įxample: Horizontal stacking of numpy arrays > import numpy as np 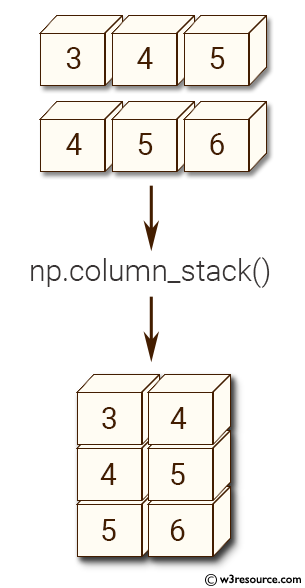
Save_image( image_array = an_image, path_without_file_name = "./output/" + manifold_learning_method + "/directions/", file_name = str( projection_direction_index) + ".The arrays must have the same shape along all but the first axis. imresize( arr = an_image, size = 500) # -> 5 times bigger reshape(( image_height, image_width))Īn_image = scipy. 
shapeįor projection_direction_index in range( n_projection_directions_to_save):Īn_image = projection_directions. N_projection_directions_to_save = projection_directions. If n_projection_directions_to_save = None: My_manifold_learning = My_kernel_supervised_PCA_UsingDirect( n_components = None, kernel_on_labels = kernel_on_labels_in_SPCA, kernel = kernel) My_manifold_learning = My_kernel_supervised_PCA_UsingDual( n_components = None, kernel_on_labels = kernel_on_labels_in_SPCA, kernel = kernel)Įlif manifold_learning_method = "kernel_SPCA_UsingDirect": My_manifold_learning = My_dual_supervised_PCA( n_components = None, kernel_on_labels = kernel_on_labels_in_SPCA)Įlif manifold_learning_method = "kernel_SPCA_UsingDual": fit_transform( X = X_train, Y = Y_train)Įlif manifold_learning_method = "dual_supervised_PCA": My_manifold_learning = My_supervised_PCA( n_components = None, kernel_on_labels = kernel_on_labels_in_SPCA)ĭata_transformed = my_manifold_learning. My_manifold_learning = My_kernel_PCA( n_components = None, kernel = kernel)Įlif manifold_learning_method = "supervised_PCA": My_manifold_learning = My_dual_PCA( n_components = None)Įlif manifold_learning_method = "kernel_PCA": get_projection_directions()Įlif manifold_learning_method = "dual_PCA": Projection_directions = my_manifold_learning. My_manifold_learning = My_PCA( n_components = None)ĭata_transformed = my_manifold_learning. # - apply manifold learning (fit + project training data + projection directions): column_stack(( X_test_notNormalized, X_class_notNormalized)) column_stack(( X_train_notNormalized, X_class_notNormalized)) X_class_notNormalized = data_notNormalized shape, 0))įor class_index in range( 1, n_classes + 1): X_train_notNormalized = data_notNormalized X_test_notNormalized = X_test_notNormalized. X_train_notNormalized = X_train_notNormalized. X_train, X_test = train_test_split( data. Img = load_image( address_image = path_dataset + "class" + str( class_index + 1) + "/" + filename) listdir( path_dataset + "class" + str( class_index + 1) + "/"): floor( image_index / 10) + 1įor filename in os. Img = load_image( address_image = path_dataset + str( image_index + 1) + ".jpg") zeros(( image_height * image_width, n_samples)) Save_image( image_array = an_image, path_without_file_name = "./Frey_dataset/images/", file_name = str( image_index) + ".png")ĭata = np. imresize( arr = an_image, size = 500) #-> 5 times bigger Scaler = StandardScaler( with_mean = True, with_std = True). # - read mat dataset and convert to csv:Ĭonvert_mat_to_csv( path_mat = path_dataset, path_to_save = path_dataset)ĭata = read_csv_file( path = path_dataset + ".csv") Path_dataset = "./Frey_dataset/frey_rawface" Which_dimensions_to_plot_inpointsAndImagesPlot = #-> list of two indices (start and end), e.g. Plot_projected_pointsAndImages_again = True OutOfSample_indices_reconstructed_images_to_save = None Indices_reconstructed_images_to_save = None OutOfSample_indices_reconstructed_images_to_save = Indices_reconstructed_images_to_save = #-> list of two indices (start and end), e.g. Save_reconstructed_outOfSample_images_again = False N_projection_directions_to_save = 10 #-> an integer >= 1, if None: save all "specified" directions when creating the python class Process_out_of_sample_all_together = True Reconstruct_using_howMany_projection_directions = None # -> an integer >= 1, if None: using all "specified" directions when creating the python class Kernel_on_labels_in_SPCA = "linear" #kernel over labels (Y) -> ‘rbf’, ‘sigmoid’, ‘polynomial’, ‘poly’, ‘linear’, ‘cosine’ -> if None, it is linear Kernel = "linear" #kernel over data (X) -> ‘rbf’, ‘sigmoid’, ‘polynomial’, ‘poly’, ‘linear’, ‘cosine’ -> if None, it is linear Manifold_learning_method = "supervised_PCA" #-> PCA, dual_PCA, kernel_PCA, supervised_PCA, dual_supervised_PCA, kernel_SPCA_UsingDual, kernel_SPCA_UsingDirect, MDS, Isomap, Laplacian_eigenmap model_selection import train_test_split #-> ĭataset = "ATT" #-> Frey, ATT, ATT_glasses preprocessing import StandardScalerįrom sklearn. From my_supervised_PCA import My_supervised_PCAįrom my_dual_supervised_PCA import My_dual_supervised_PCAįrom my_kernel_supervised_PCA_UsingDual import My_kernel_supervised_PCA_UsingDualįrom my_kernel_supervised_PCA_UsingDirect import My_kernel_supervised_PCA_UsingDirectįrom sklearn.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
